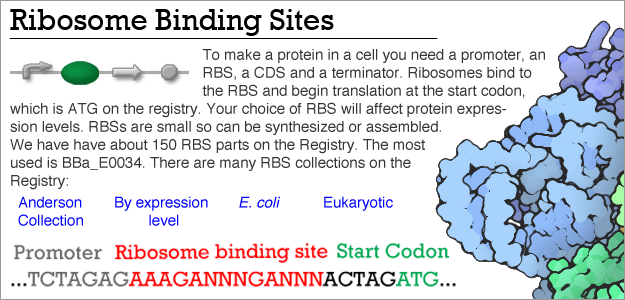
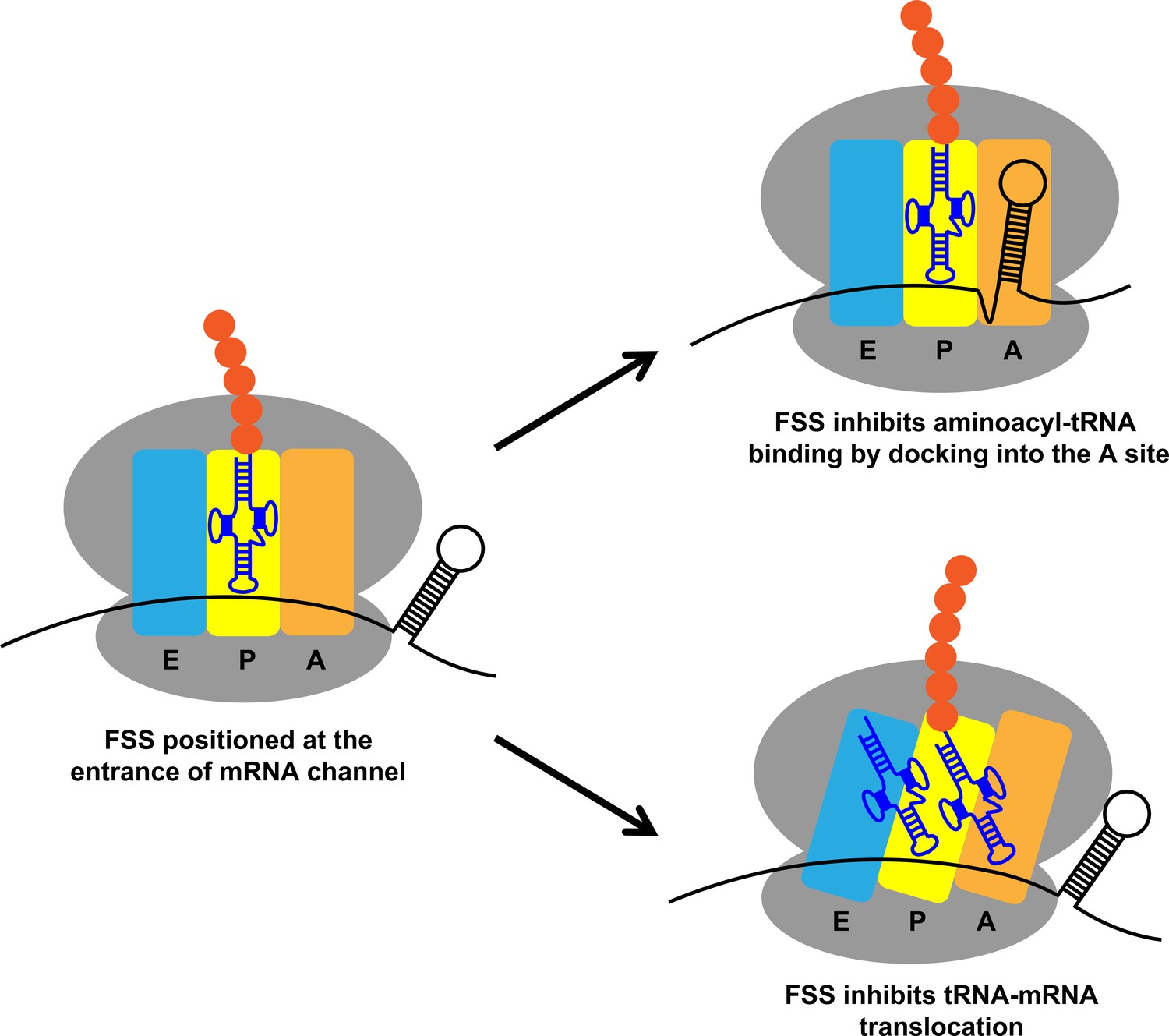
Influence of the spacer region between the Shine–Dalgarno box and the start codon for fine‐tuning of the translation efficiency in Escherichia coli - Komarova - 2020 - Microbial Biotechnology - Wiley Online Library
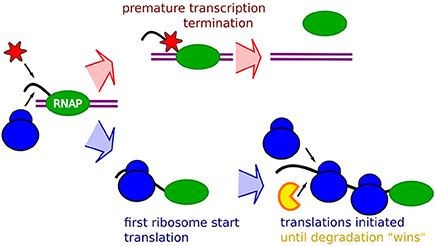
Frontiers | Occlusion of the Ribosome Binding Site Connects the Translational Initiation Frequency, mRNA Stability and Premature Transcription Termination

i) DNA templates containing a T7 promoter (1), ribosomal binding site... | Download Scientific Diagram
GitHub - hsalis/Ribosome-Binding-Site-Calculator-v1.0: The Ribosome Binding Site (RBS) Calculator can predict the translation initiation rate of a protein coding sequence in bacteria and design synthetic RBS sequences to rationally control the translation

Sequence logos of putative promoters, ribosome-binding sites (RBS) and... | Download Scientific Diagram

1 The ribosome binding site and 5 0 protein coding sequence control an... | Download Scientific Diagram
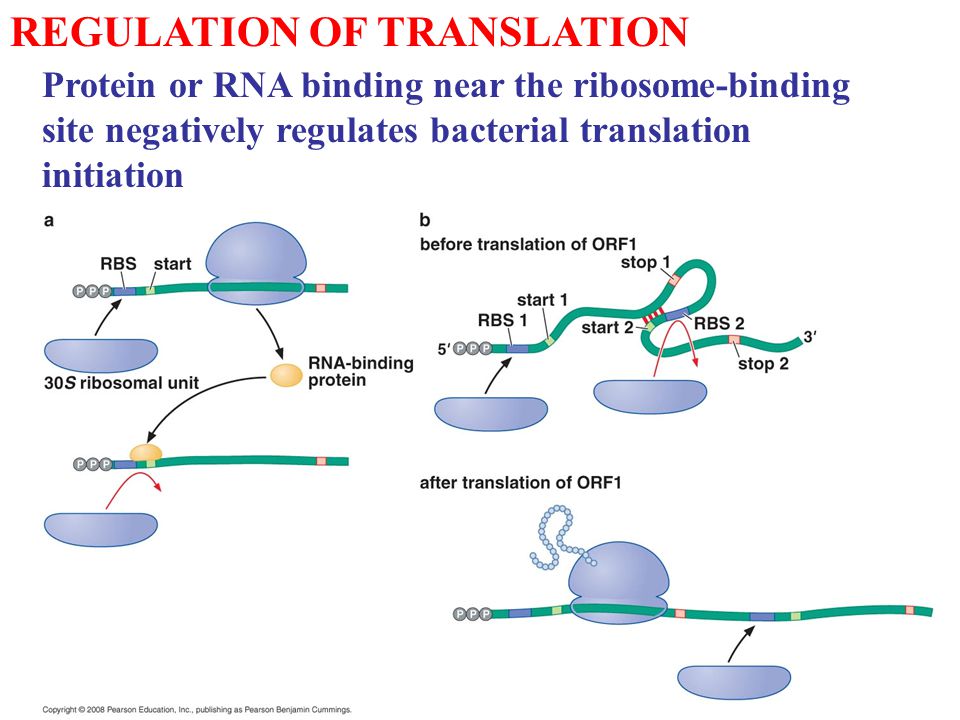
REGULATION OF TRANSLATION Protein or RNA binding near the ribosome-binding site negatively regulates bacterial translation initiation. - ppt download

Deciphering the Rules of Ribosome Binding Site Differentiation in Context Dependence | ACS Synthetic Biology
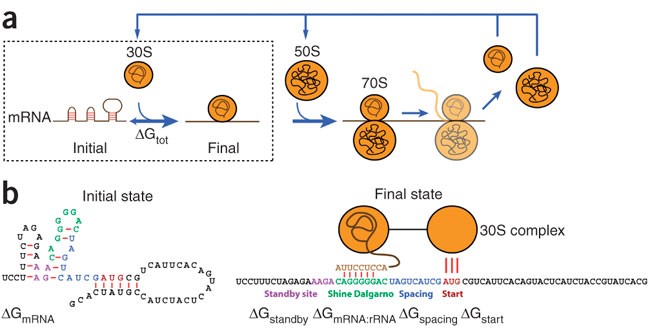
Automated design of synthetic ribosome binding sites to control protein expression | Nature Biotechnology

rClone: A Synthetic Biology Tool That Enables the Research of Bacterial Translation — Journal of Young Investigators

Deciphering the Rules of Ribosome Binding Site Differentiation in Context Dependence | ACS Synthetic Biology
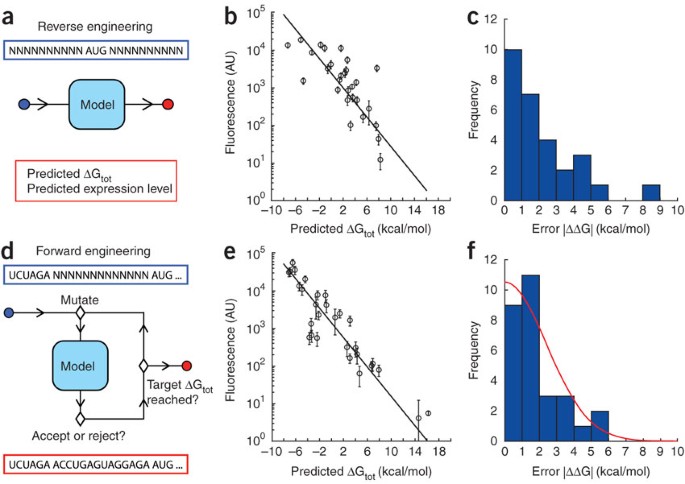
Automated design of synthetic ribosome binding sites to control protein expression | Nature Biotechnology

SciELO - Brasil - Ribosome binding site recognition using neural networks Ribosome binding site recognition using neural networks

The context of the ribosome binding site in mRNAs defines specificity of action of kasugamycin, an inhibitor of translation initiation | PNAS


![PDF] Classification of Escherichia coli K-12 ribosome binding sites | Semantic Scholar PDF] Classification of Escherichia coli K-12 ribosome binding sites | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/4af616123bce83b1f5dff449961c1df697c8d771/2-Figure2-1.png)